Difference between revisions of "Goals"
| Line 5: | Line 5: | ||
'''Specific Aim 2''': Extract gene/protein mentions and their variants from the clinical narrative | '''Specific Aim 2''': Extract gene/protein mentions and their variants from the clinical narrative | ||
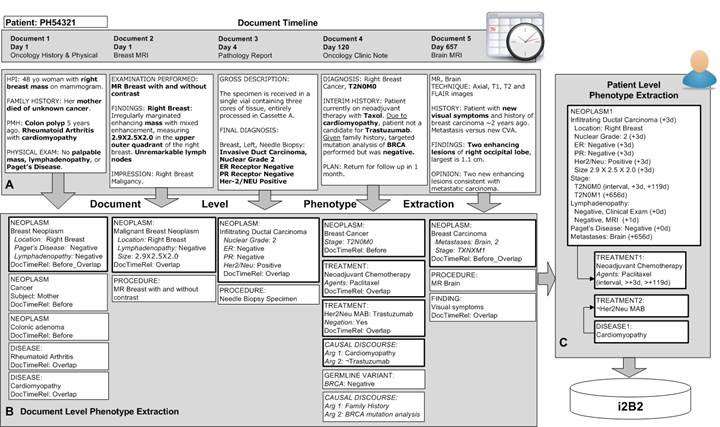
| − | '''Specific Aim 3''': Create longitudinal representation of disease process and its resolution. Link phenotypes, treatments and outcomes in temporal associations to create a longitudinal abstraction of the disease | + | '''Specific Aim 3''': Create longitudinal representation of disease process and its resolution. Link phenotypes, treatments and outcomes in temporal associations to create a longitudinal abstraction of the disease. |
| + | |||
[[File:Aim 3.jpg]] | [[File:Aim 3.jpg]] | ||
| + | |||
'''Specific Aim 4''': Extract discourses containing explanations, speculations, and hypotheses, to support explorations of causality | '''Specific Aim 4''': Extract discourses containing explanations, speculations, and hypotheses, to support explorations of causality | ||
Revision as of 09:52, 16 August 2016
Our first four development specific aims significantly extend the capability of our current software, focusing on challenging problems in biomedical information extraction. These aims support the development and evaluation of novel methods for cancer deep phenotype extraction:
Specific Aim 1: Develop methods for extracting phenotypic profiles. Extract patient’s deep phenotypes, and their attributes such as general modifiers (negation, uncertainty, subject) and cancer specific characteristics (e.g. grade, invasion, lymph node involvement, metastasis, size, stage)
Specific Aim 2: Extract gene/protein mentions and their variants from the clinical narrative
Specific Aim 3: Create longitudinal representation of disease process and its resolution. Link phenotypes, treatments and outcomes in temporal associations to create a longitudinal abstraction of the disease.
Specific Aim 4: Extract discourses containing explanations, speculations, and hypotheses, to support explorations of causality
Our last two implementation specific aims focus on the design of the software to support the cancer research community, ensuring the usability and utility of our software. These aims support the design, dissemination and sharing of the products of this work to maximize impact on cancer research:
Specific Aim 5: Design and implement a computational platform for deep phenotype discovery and analytics for translational investigators, including integrative visual analytics.
Specific Aim 6: Advance translational research in driving cancer biology research projects in breast cancer, ovarian cancer, and melanoma. Include research community throughout the design of the platform and its evaluation. Disseminate freely available software.
Impact: The proposed work will produce novel methods for extracting detailed phenotype information directly from the EMR, the major source of such data for patients with cancer. Extracted phenotypes will be used in three ongoing translational studies with a precision medicine focus. Dissemination of the software will enhance the ability of cancer researchers to abstract meaningful clinical data for translational research. If successful, systematic capture and representation of these phenotypes from EMR data could later be used to drive clinical genomic decision support.